There are currently two major pressures on crystallographic computing:
- Increased automation to increase throughput in line with genomics applications.
- Increased data complexity, as data from more sources is combined and carried through the whole calculation to solve more difficult problems.
The Clipper project is an initiative to address these pressures.
The aim of the project is to produce a set of object-oriented libraries for the organisation of crystallographic data and the performance of crystallographic computation. The libraries are designed as a framework for new crystallographic software, which will allow the full power of modern programming techniques to be exploited by the developer. This will lead to greater functionality from simpler code which will be easier to develop and debug.
Object oriented programming
The evolution of high level programing may be very imprecisely caricatured as follows:
- Early high-level languages (Fortran): There is no structure to data or code. Variables are not grouped in any way or specially associated with any code.
- Structured programming (C, Pascal, etc): Data structures are introduced, grouping related variables into compound objects, which can completely specify the state of some object.
- Object-oriented programming (Smalltalk, C++, etc): Data structures may now contain code, and now describe not only the state of the object, but also its behavior and interactions. (In the general case, all code becomes part of an object.)
Clipper is object-oriented. The main benefit of this approach is that code becomes much more reusable, since objects are self-contained, and may be reused, rewritten, or replaced without affecting other code. Additionally, the organisation of the code and data is generally much clearer.
Clipper objects
Clipper defines a wide range of objects. These fall into a number of groups, including:
- Crystallographic objects: : Cell, Spacegroup, Metric, R/T operator, etc.
- Data objects: : Reflection data, Crystallographic map, Non-crystallographic map, Coordinate model.
- Method objects: : FFT arrays, Resolution functions, Conversion objects, Import/export objects
The coordinate object is a substantial task in itself and is addressed by an interface to the CCP4 'MMDB' library, developed by Dr Eugene Krissinel.
Some of the objects will be discussed in more detail:
Crystallographic objects:
These implements the fundamental properties of a crystal.
- The Cell object:
This object describes a unit cell. It holds the cell parameters, and derived information including coordinate conversion matrices and metrics. Any cell object may be used to convert coordinates between orthogonal and fractional forms, and calculate distances in real space and resolutions in reciprocal space.
- The Spacegroup object:
- This object describes a spacegroup, using information from the 'cctbx' library of Dr Ralph Grosse-Kunstleve. It can be use to generate symmetry coordinates and reflections, phase shifts, centricity, asymmetric units, and so on.
Data objects:
These hold actual data. They are written as templates which can hold whatever type of data the developer requires.
- The reflection data object:
It is commonly necessary to store several related items of reflection data. Therefore this object is split into two parts; a parent object which holds a list of Miller indices and related data, and then several data objects which hold the actual data associated with each Miller index. The data objects can hold data of arbitrary types: these types will usually consist of several values. For example, a structure factor magnitude and its variance, or all four Hendrickson-Lattman coefficients, are usually held in a single data object.
To the user, the data appears to cover the whole of reciprocal space, however in practice only an asymmetric unit is stored. Data is transformed about reciprocal spaces as required. When a new data type is defined, its behavior under transformation is also be defined so that this mapping can be performed.
- The crystallographic map object:
This object also implements crystallographic symmetry, and also cell repeat, in a manner which is transparent to the user. It may also hold arbitrary data types: common examples would include bits, real values, complex values, or orthogonal or fractional gradients.
- The non-crystallographic map object:
- This object is used for map data which does not have symmetry or cell repeat, for example an NCS averaging mask.
Method objects:
These are used to provide additional functionality commonly required in crystallographic calculations. Examples include:
- FFT map:
This object holds data which may be represented in either real or reciprocal space. The data may be accessed in either form, and may be transformed between spaces as required.
- The resolution function evaluator:
This object creates an arbitrary function of position in reciprocal space, by optimising the parameters of some basis function in order to minimise some target function. This is an extreme generalisation of the idea of 'resolution bins', and can be used for anything from <|F|2>s to sigma-a and beyond.
- Import/export objects:
- The only import/export objects implemented so far are the MTZfile and MAPfile objects. These stand as proxies for an external file, and can transfer data between the Clipper objects and the file.
Scripting
Automation of crystallographic tasks depends on being able to communicate between successive tasks, and by being able to execute control code to activate tasks and make protocol decisions. This functionality is provided through a scripting interface. A Python interface will be provided through the boost.python library. It is possible that interfaces to C and a range of scripting languages will be provided through other means. All the data and the full functionality of the methods will be available from the scripting layer, allowing full automation and full communication between tasks.
Eventually the individual programs should disappear, rather exposing their functionality directly to the scripting layer. At the same time, data will have to move from traditional files into a database, so that each task has immediate access to all the information currently available.
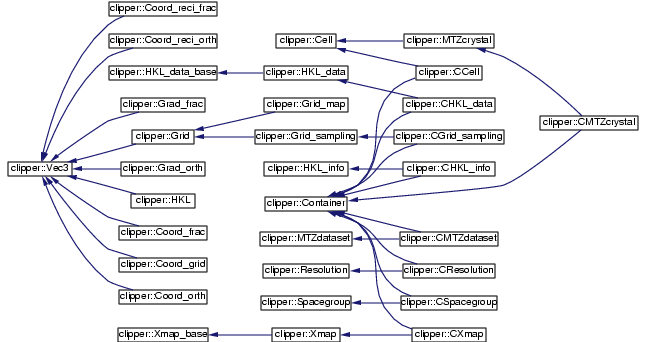
Partial class hierarchy (as of 1/2/2001)
Acknowledgments:
I would like to thank Ralph Grosse-Kunstleve, Airlie McCoy, Eugene Krissinel, Jan Zelinka and the CCP4 staff for their many and varied contributions to this effort.
Clipper stands for 'Cross-crystal Likelihood Phase Probability Estimation and Refinement', which is what I hope to use it for.
References:
- 'Clipper' code and documentation (K. Cowtan): http://www.ysbl.york.ac.uk/~cowtan/clipper/clipper.html
- 'mmdb' coordinate library (E. Krissinel): http://www.ebi.ac.uk/~keb/cldoc/
- 'cctbx' crystallography toolbox (R. Grosse-Kunstleve): http://cctbx.sourceforge.net
See the following pages in the Clipper documentation for further information:
- Class Overview (brief guide to classes)
- Conventions (coding style)
- Coordinates and related types (Coodinate types)
- Developing using Clipper (initial example)
- Developing using Reflection Data (reflection data handling and examples)
- Developing using Crystallographic Maps (map handling and examples)
- Developing using Atomic Models (model handling)
- Where to Look for help on specific tasks. (thematic index)
- Installation (and dependencies)
 1.7.4
1.7.4